 Remoscope: a label-free imaging cytometer for malaria diagnostics... Trans R Soc Trop Med Hyg 2025
|
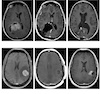 IL-6 underlies microenvironment immunosuppression and resistance to therapy in glioblastoma bioRxiv, 2025
|
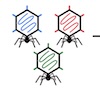 A Trypanosoma cruzi Trans-Sialidase Peptide Demonstrates High Serological Prevalence Among Infected Populations Across Endemic Regions of Latin America medRxiv, 2025
|
 Proteomic profiling of the local and systemic immune response to pediatric respiratory viral infections American Society for Microbiology, 2025
|
 Endogenous antigens shape the transcriptome and TCR repertoire in an autoimmune arthritis model The Journal of Clinical Investigation, 2024
|
 Unveiling the proteome-wide autoreactome enables enhanced evaluation of emerging CAR T cell therapies in autoimmunity J Clin Invest 2024
|
 Total Syntheses of Cyclomarin and Metamarin Natural Products Organic Letters, 2024
|
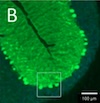 Anti-RGS8 paraneoplastic cerebellar ataxia is preferentially associated with a particular subtype of Hodgkin's lymphoma Springer Nature, 2024
|
 Microbial dynamics and pulmonary immune responses in COVID-19 secondary bacterial pneumonia Nature Portfolio, 2024
|
 Climate, demography, immunology, and virology combine to drive two decades of dengue virus dynamics in Cambodia PNAS, 2024
|
 Molecular mimicry in multisystem inflammatory syndrome in children Nature Portfolio, 2024
|
 Phage Immunoprecipitation-Sequencing Reveals CDHR5 Autoantibodies in Select Patients With Interstitial Lung Disease American College of Rheumatology, 2024
|
|  Here we introduce HMMSplicer, an accurate and efficient algorithm for discovering canonical and non-canonical splice junctions in short read datasets. HMMSplicer identifies more splice junctions than currently available algorithms when tested on publicly available A. thaliana, P. falciparum, and H. sapiens datasets without a reduction in specificity. HMMSplicer was found to perform especially well in compact genomes and on genes with low expression levels, alternative splice isoforms, or non-canonical splice junctions. Because HHMSplicer does not rely on pre-built gene models, the products of inexact splicing are also detected. In addition, HMMSplicer provides a score for every predicted junction allowing the user to set a threshold to tune false positive rates depending on the needs of the experiment. HMMSplicer is implemented in Python. Code and documentation are freely available at the link below. Download HMMSplicer! (Updated Nov 25th 2010 - Version: 0.9.5) |
|
